Supervisor
Jake Harris
Brief Summary
Epigenetic marks provide the cell with information about the underlying DNA. This project aims to understand how these marks are decoded by the cell.
Importance of Research
Gene regulation is fundamental to life, and in eukaryotes nucleosomes act as a first barrier to transcription. Therefore, histone marks critically affect a genes ability to respond appropriately to biotic and abiotic stresses. Knowledge of how they these marks function opens the door to engineering plants with enhanced resilience.
Project Summary
Regulating access to DNA is a primary means by which gene expression states are controlled. Epigenetic marks can directly influence transcription by affecting DNA accessibility, and by serving as molecular signposts for ‘reader’ proteins that can recruit or repel transcriptional machinery. Over the past 20 years, we have learned where these marks are in the genome, but our understanding of how these marks are read and perceived by the cell lags far behind. Identifying these reader proteins is an essential step in understanding the links between particular epigenetic states and their functions.
What will the successful application do?
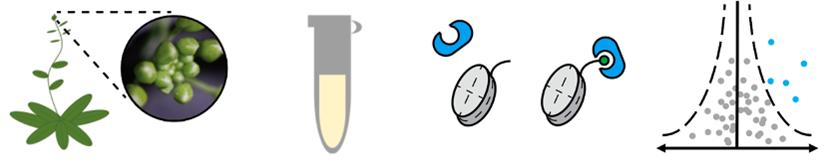
The first stage of this project will be to identify the readers of key histone marks. The work will involve a quantitative proteomics based ‘molecular fishing’ approach - performed in collaboration with world experts in Germany. In the second phase of the work, readers proteins will be assessed using genomics (ChIP-seq/CUT&Tag and RNA-seq), transcriptional reporter assays, and biochemical approaches, to gain a mechanistic understanding of the role and function of key histone reader proteins.
Training Provided
The student will gain expertise in a range standard molecular biology techniques (cloning, genotyping, qRT-PCR, western blots). In addition the student will learn biochemical (protein purifications, binding / enzymatic assays) and genomic (both wet lab and computational) approaches.
References
- Scheid R, Chen J, Zhong X. Biological role and mechanism of chromatin readers in plants. Curr Opin Plant Biol. (2021) Jun;61:102008. doi: 10.1016/j.pbi.2021.102008. Epub 2021 Feb 10. PMID: 33581373; PMCID: PMC8222062.
- Harris CJ, Scheibe M, Wongpalee SP, Liu W, Cornett EM, Vaughan RM, Li X, Chen W, Xue Y, Zhong Z, Yen L, Barshop WD, Rayatpisheh S, Gallego-Bartolome J, Groth M, Wang Z, Wohlschlegel JA, Du J, Rothbart SB, Butter F, Jacobsen SE. A DNA methylation reader complex that enhances gene transcription. Science. 2018 Dec 7;362(6419):1182-1186. doi: 10.1126/science.aar7854. PMID: 30523112; PMCID: PMC6353633.